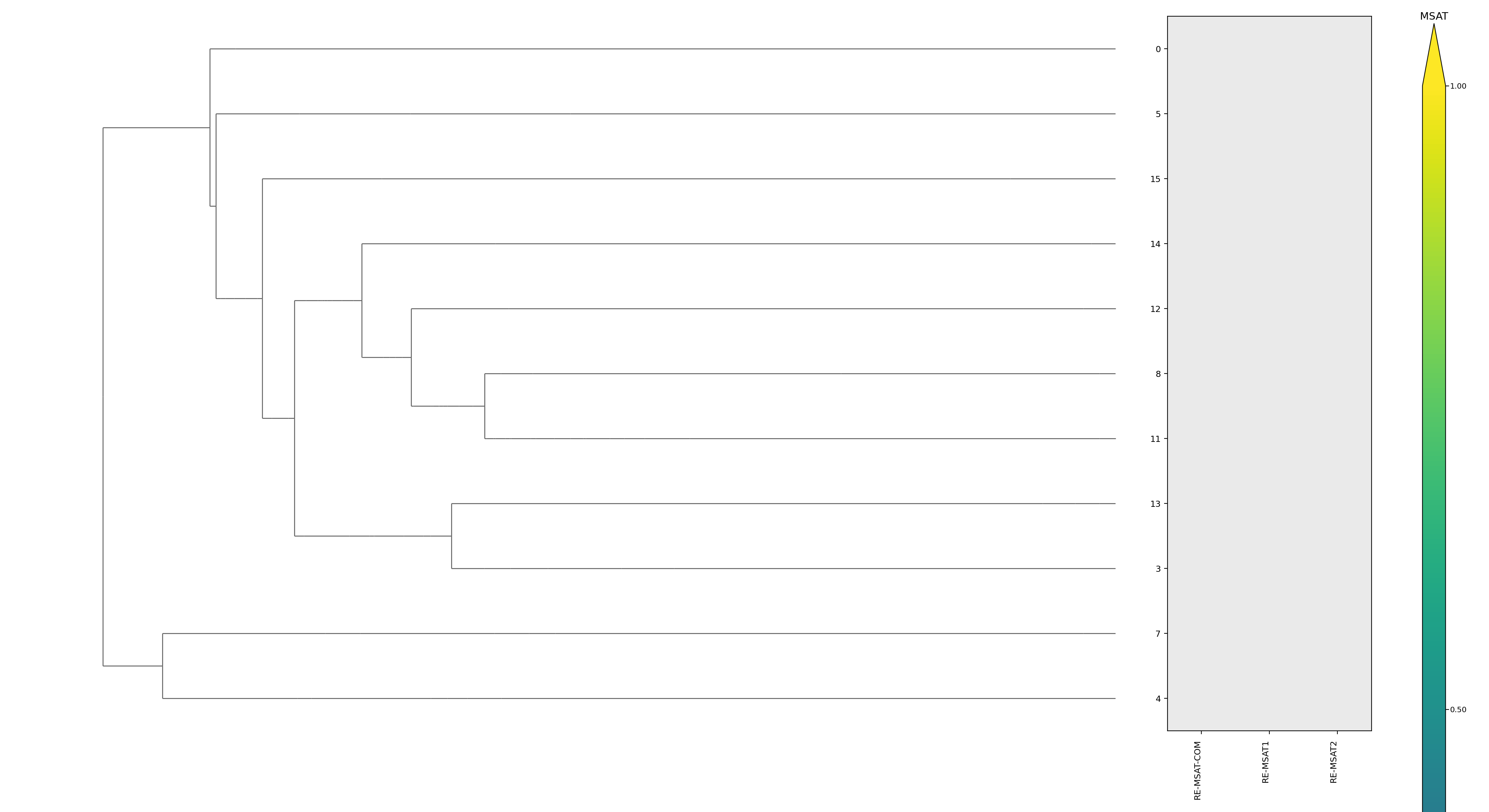
| cluster | n | neff | AF2_TM1 | AF2_TM2 | AF3_TM1 | AF3_TM2 | RE-MSAT-COM | RE-MSAT1 | RE-MSAT2 | ESM_SEQIDS | ESM_TM1_LIST | ESM_TM2_LIST |
|---|
| Deep | - | - | 0.78 | 0.88 | 0.78 | 0.89 | 0.24 | 0.3 | 0.18 | - | - | - |
| 000 | - | - | 0.74 | 0.75 | 0.77 | 0.71 | 0.03 | 0.02 | 0.03 | 008; 008; 001; 007; 007; 001; 002; 002; 003; 003 | 0.74; 0.74; 0.73; 0.73; 0.73; 0.73; 0.72; 0.72; 0.70; 0.69 | 0.83; 0.83; 0.76; 0.83; 0.83; 0.76; 0.82; 0.82; 0.79; 0.80 |
| 001 | - | - | 0.63 | 0.73 | 0.65 | 0.79 | 0.0 | 0.0 | 0.0 | 001; 002; 006; 004; 003; 002; 001; 006; 004; 003 | 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74 | 0.85; 0.80; 0.85; 0.78; 0.85; 0.80; 0.85; 0.85; 0.78; 0.85 |
| 002 | - | - | 0.68 | 0.63 | 0.68 | 0.63 | 0.0 | 0.0 | 0.0 | 009; 009; 004; 004; 002; 002; 001; 001; 005; 005 | 0.72; 0.72; 0.72; 0.72; 0.68; 0.68; 0.67; 0.67; 0.58; 0.57 | 0.69; 0.69; 0.70; 0.70; 0.66; 0.66; 0.63; 0.63; 0.56; 0.56 |
| 003 | - | - | 0.63 | 0.73 | 0.65 | 0.79 | 0.09 | 0.08 | 0.13 | 001; 002; 006; 004; 003; 002; 001; 006; 004; 003 | 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74; 0.74 | 0.85; 0.80; 0.85; 0.78; 0.85; 0.80; 0.85; 0.85; 0.78; 0.85 |
| 004 | - | - | 0.68 | 0.63 | 0.68 | 0.63 | 0.0 | 0.0 | 0.0 | 009; 009; 004; 004; 002; 002; 001; 001; 005; 005 | 0.72; 0.72; 0.72; 0.72; 0.68; 0.68; 0.67; 0.67; 0.58; 0.57 | 0.69; 0.69; 0.70; 0.70; 0.66; 0.66; 0.63; 0.63; 0.56; 0.56 |
| 005 | - | - | 0.74 | 0.8 | 0.73 | 0.79 | 0.0 | 0.0 | 0.01 | 001; 008; 008; 001; 009; 009; 002; 002; 003; 003 | 0.75; 0.75; 0.75; 0.75; 0.73; 0.73; 0.72; 0.72; 0.71; 0.71 | 0.80; 0.84; 0.84; 0.80; 0.86; 0.86; 0.81; 0.81; 0.83; 0.83 |
| 006 | - | - | 0.69 | 0.82 | 0.62 | 0.76 | 0.0 | 0.0 | 0.0 | 002; 009; 004; 004; 009; 002; 001; 007; 006; 003 | 0.76; 0.76; 0.76; 0.76; 0.76; 0.76; 0.75; 0.75; 0.75; 0.75 | 0.87; 0.78; 0.78; 0.78; 0.77; 0.87; 0.88; 0.86; 0.86; 0.87 |
| 007 | - | - | 0.42 | 0.56 | 0.38 | 0.53 | 0.0 | 0.01 | 0.0 | 002; 002; 006; 006; 003; 003; 008; 008; 001; 001 | 0.74; 0.74; 0.73; 0.73; 0.68; 0.68; 0.45; 0.45; 0.44; 0.44 | 0.80; 0.80; 0.80; 0.80; 0.76; 0.76; 0.59; 0.59; 0.58; 0.58 |
| 008 | - | - | 0.75 | 0.85 | 0.74 | 0.86 | 0.0 | 0.0 | 0.0 | 001; 002; 002; 001; 007; 007; 008; 008; 003; 003 | 0.76; 0.76; 0.76; 0.76; 0.75; 0.75; 0.72; 0.72; 0.69; 0.69 | 0.87; 0.87; 0.87; 0.87; 0.87; 0.87; 0.85; 0.85; 0.69; 0.69 |
| 009 | - | - | 0.75 | 0.85 | 0.76 | 0.86 | 0.0 | 0.01 | 0.0 | 001; 007; 007; 001; 006; 006; 002; 002; 005; 005 | 0.76; 0.76; 0.76; 0.76; 0.74; 0.74; 0.72; 0.72; 0.71; 0.71 | 0.86; 0.84; 0.84; 0.86; 0.84; 0.84; 0.76; 0.76; 0.82; 0.82 |
| 010 | - | - | 0.76 | 0.87 | 0.76 | 0.86 | 0.03 | 0.04 | 0.08 | 001; 001; 008; 008; 006; 007; 006; 007; 005; 005 | 0.74; 0.74; 0.73; 0.73; 0.72; 0.72; 0.72; 0.72; 0.70; 0.70 | 0.86; 0.86; 0.83; 0.83; 0.84; 0.83; 0.84; 0.83; 0.69; 0.69 |
| 011 | - | - | 0.69 | 0.83 | 0.62 | 0.76 | - | - | - | 002; 009; 004; 004; 009; 002; 001; 007; 006; 003 | 0.76; 0.76; 0.76; 0.76; 0.76; 0.76; 0.75; 0.75; 0.75; 0.75 | 0.87; 0.78; 0.78; 0.78; 0.77; 0.87; 0.88; 0.86; 0.86; 0.87 |
| 012 | - | - | 0.76 | 0.87 | 0.76 | 0.86 | - | - | - | 009; 009; 003; 003; 002; 002; 001; 001; 007; 007 | 0.77; 0.77; 0.76; 0.76; 0.76; 0.76; 0.75; 0.75; 0.74; 0.74 | 0.86; 0.86; 0.87; 0.87; 0.86; 0.86; 0.86; 0.86; 0.85; 0.85 |
| 013 | - | - | 0.75 | 0.85 | 0.76 | 0.85 | - | - | - | 001; 005; 005; 001; 006; 006; 007; 007; 002; 010 | 0.75; 0.75; 0.75; 0.75; 0.72; 0.72; 0.71; 0.71; 0.57; 0.54 | 0.86; 0.86; 0.86; 0.86; 0.83; 0.83; 0.81; 0.81; 0.63; 0.58 |
| 014 | - | - | 0.75 | 0.85 | 0.76 | 0.86 | - | - | - | 001; 007; 007; 001; 006; 006; 002; 002; 005; 005 | 0.76; 0.76; 0.76; 0.76; 0.74; 0.74; 0.72; 0.72; 0.71; 0.71 | 0.86; 0.84; 0.84; 0.86; 0.84; 0.84; 0.76; 0.76; 0.82; 0.82 |
| 015 | - | - | 0.76 | 0.86 | 0.76 | 0.86 | - | - | - | 001; 001; 008; 008; 006; 007; 006; 007; 005; 005 | 0.74; 0.74; 0.73; 0.73; 0.72; 0.72; 0.72; 0.72; 0.70; 0.70 | 0.86; 0.86; 0.83; 0.83; 0.84; 0.83; 0.84; 0.83; 0.69; 0.69 |